Authentic Research Experience and “Big Data” Analysis in the Classroom: Maize Response to Abiotic Stress
Abstract
Integration of inquiry-based approaches into curriculum is transforming the way science is taught and studied in undergraduate classrooms. Incorporating quantitative reasoning and mathematical skills into authentic biology undergraduate research projects has been shown to benefit students in developing various skills necessary for future scientists and to attract students to science, technology, engineering, and mathematics disciplines. While large-scale data analysis became an essential part of modern biological research, students have few opportunities to engage in analysis of large biological data sets. RNA-seq analysis, a tool that allows precise measurement of the level of gene expression for all genes in a genome, revolutionized molecular biology and provides ample opportunities for engaging students in authentic research. We developed, implemented, and assessed a series of authentic research laboratory exercises incorporating a large data RNA-seq analysis into an introductory undergraduate classroom. Our laboratory series is focused on analyzing gene expression changes in response to abiotic stress in maize seedlings; however, it could be easily adapted to the analysis of any other biological system with available RNA-seq data. Objective and subjective assessment of student learning demonstrated gains in understanding important biological concepts and in skills related to the process of science.
INTRODUCTION
Integration of inquiry-based approaches into curriculum is transforming the way science is taught and studied in undergraduate classrooms (National Research Council [NRC], 2003; American Association for the Advancement of Science [AAAS], 2011). Reviews of novel curricular approaches in undergraduate science courses suggest that teaching practices are consistently changing in the direction of inquiry, with greater than 80% of the institutions using inquiry-based approaches, especially in their laboratory courses (Sundberg et al., 2005; Ruiz-Primo et al., 2011). Authentic undergraduate research experiences have been repeatedly shown to benefit students in a variety of ways, leading them to learn to think like a scientist, find research exciting, and pursue graduate education or careers in science (Lopatto et al., 2008, 2014; Thiry and Laursen, 2011). Moving from guided-inquiry toward research-based laboratory approaches in introductory undergraduate science courses has been shown to keep students interested in science and to prepare them for future careers (Weaver et al., 2008). Several national projects working to advance course-based research experiences have been successful in providing students with high-impact learning experiences in biology research (Hanauer et al., 2006; Campbell et al., 2007; Ditty et al., 2010; Laursen et al., 2010; Shaffer et al., 2010). The Course-based Undergraduate Research Network (CUREnet) provides support and helps foster collaboration among faculty interested in incorporating research experiences into their classrooms (Auchincloss et al., 2014). In addition, tools have been developed that allow assessment of student learning as a result of research experiences in individual courses and comparison of individual courses with other courses with embedded research experiences (Lopatto et al., 2008, 2014). Despite these promising trends, a recent survey of inquiry-based teaching in biology laboratory courses found that greater than 75% of inquiry-based laboratory studies were guided-inquiry exercises rather than research experiences embedded in a course (Beck et al., 2014; Spell et al., 2014). Research experiences provide students with a higher degree of independence in defining the research question and approaches to data analysis. Also, unlike in guided-inquiry exercises, the outcomes of the research are not known and hold the potential for generating new scientific knowledge (Weaver et al., 2008). Greater than 75% of research experiences embedded in a course were targeted toward upper-level biology students rather than students in introductory courses (Beck et al., 2014). Thus, development and integration of research experiences into introductory biology courses remains an important target in teaching science at the undergraduate level.
The recent explosion of big data in various fields of science, including biology, has led to high demand for integration of big data analysis with computational and quantitative thinking into the skill set required of graduates. “Data scientists” have been described as having the “sexiest job of the 21st century” (Davenport and Patil, 2012). However, many biology graduates lack sufficient skills in mathematical and quantitative reasoning, analysis and visualization of big data, and cross-disciplinary approaches required to solve complex biological problems (Feser et al., 2013; Magana et al., 2014). Hence, it is vital to develop effective educational approaches and strategies for improving students’ mathematical reasoning in solving biological problems (Hester et al., 2014), including students’ skills in analyzing large biological data sets. Approaches to integrating bioinformatics analysis to the curriculum target primarily DNA sequence analysis through BLAST and other National Center for Biotechnology Information (NCBI) tools and primary literature research through PubMed, and are focused primarily on analysis of single genes/proteins and gene families as opposed to large-scale genomics data interpretations (Magana et al., 2014).
Most successful implementations of bioinformatics and genomics course modules are built around important biological concepts and are aimed at enhancing student understanding of these concepts (Magana et al., 2014). One of the central biological concepts is the regulation of gene expression in response to changes in the environment or through various developmental stages. Large-scale gene expression analysis using microarrays was recently implemented in several different projects and was proven to be a powerful education tool targeting important and widely applicable biological concepts (Campbell et al., 2007). RNA-seq analysis, a technique used to quantify the amount of RNA transcribed from each gene of an organism, has become a prevalent method for researchers investigating regulation of gene expression in a variety of biological systems. It could serve as a great tool to help students understand the principles of gene expression regulation. Even though producing RNA-seq data sets remains a relatively expensive endeavor for many institutions, numerous RNA-seq data sets collected in various systems are freely available through the Sequence Read Archive (SRA) of NCBI and could be mined to answer a variety of biologically relevant questions (www.ncbi.nlm.nih.gov/Traces/sra). Most of the bioinformatics packages researchers use to analyze RNA-seq data are also openly available and could be used by undergraduate student researchers. The availability of nearly unlimited RNA-seq data and access to powerful bioinformatics analyses from shared servers offer students the opportunity to develop into scientists while enrolled in undergraduate biology courses (Micklos et al., 2011). However, integration of RNA-seq analysis into the undergraduate curriculum is complicated by the steep learning curve the educators themselves encounter. The Genome Consortium for Active Teaching using Next-Generation Sequencing and the DNA Learning Center offer workshops that aim at helping educators gain necessary experience in the RNA-seq analysis and develop educational tools for their students (Buonaccorsi et al., 2011, 2014). The DNA Learning Center in collaboration with iPlant (Goff et al., 2011) developed a Green Line of the DNA Subway, a tool aimed at providing students with the opportunities to conduct research-grade RNA-seq analysis (http://dnasubway.iplantcollaborative.org). However, educational resources on RNA-seq analysis, including instructional materials and data analysis protocols that could be readily integrated into the classroom, are limited. Development and assessment of such resources is required to fully harness the potential of using RNA-seq analysis in the undergraduate classroom.
We present here a series of authentic research laboratory exercises incorporating a large data RNA-seq analysis into an introductory undergraduate classroom. During this laboratory module, students work on a real research project, analyzing novel data and potentially contributing to a pool of scientific knowledge. Our laboratory series is focused on analyzing gene expression changes in response to abiotic stress in maize seedlings. However, it could be easily adapted to any other RNA-seq data set. Objective and subjective assessment of student learning demonstrated gains in understanding important biological concepts and in skills related to the process of science.
METHODS
Learning Objectives and Outcomes
After completing the lab module, the students should be able to 1) explain the concepts of gene expression and transcriptional response of organisms to stress; 2) discuss the principles of RNA-seq data analysis; 3) ask scientific questions relevant to RNA-seq data analysis and identify approaches to answer these questions; 4) perform basic RNA-seq data analysis using the Green Line of the DNA Subway and DE-Seq of the R software package to assess the quality of the data and to identify genes differentially expressed between two samples; and 5) construct several types of graphs to visualize RNA-seq data.
Course Description and Student Demographics
The laboratory series on understanding plant response to abiotic stress using RNA-seq analysis was implemented as three 3-h laboratory periods (see Table 1, Figure 1, and the Supplemental Material for the details). It was conducted in 2014 during weeks 11–13 of Principles of Genetics, an introductory sophomore-level genetics course with 85 students, and in 2015 during weeks 10–12 of Applied Biotechnology, an upper-level elective course with eight students. The laboratory series on understanding transcriptional response to abiotic stress was conducted as a part of the lab component of both courses. Students worked in groups of two to four. In addition, six undergraduate research students working during Summer 2014 and Spring 2015 completed this module as part of their training for RNA-seq data analysis. Overall, 38% of the student participants were male; 17% of the students self-identified as African American, Asian American (Hmong), Hispanic, or multiracial, while 83% of the students self-identified as white. Principles of Genetics is a required course for students majoring in biology (∼60% of the students) and students majoring in exercise science (∼30% of the students), as well as for students of other majors seeking a forensic science certificate. Students majoring in biology usually take this course after taking ecology and evolution and physiology courses, although it is not unusual for the students to take genetics and ecology and evolution simultaneously. Genetics is the first course of the biology sequence in which genetics concepts, including gene expression, are covered. For many students who are not majoring in biology, genetics is their first biology course. Therefore, Principles of Genetics is considered an introductory biology course in our program. All of the Applied Biotechnology students and research students were biology majors. In all of the instances of the course implementation, most students (96%) did not have prior experience with large-scale data analysis, and none of the students had previous experience with RNA-seq data analysis, as was assessed using a precourse survey (Supplemental Material). The overwhelming majority of the students had no experience in using R or other computational approaches to data analysis and data visualization (Figure 2). When understanding of the concepts of gene expression regulation was assessed in the beginning of the course, the scores of individual students consistently fluctuated at around 25–35% (Tables 2 and 3). In Principles of Genetics, the concepts of gene expression regulation were covered in lectures/course discussions before the laboratory series on RNA-seq data analysis, and the posttest therefore likely reflects the learning gains due to both lecture and lab portions of the course. Students in Applied Biotechnology and research students scored significantly higher on the pretest compared with students in Principles of Genetics, likely suggesting the level of knowledge retained from the previously taken course in genetics that did not include RNA-seq data analysis. In Applied Biotechnology, the concepts of gene expression regulation were only discussed during the lab.
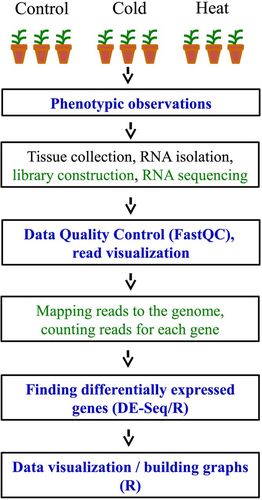
Figure 1. The flow of an RNA-seq experiment. The steps shown in blue were performed by students. Students completed learning exercises only for the steps shown in green.
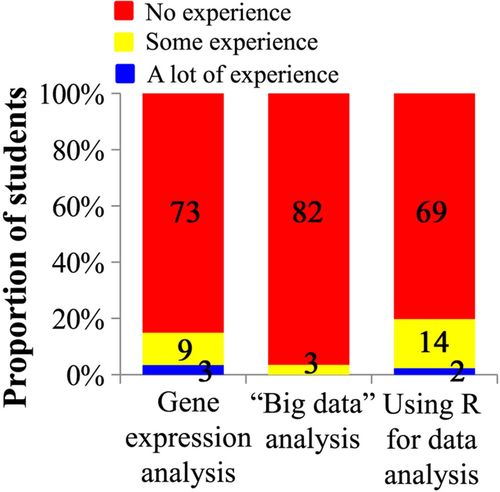
Figure 2. Students’ prior exposure to the data analysis approaches used in the lab. During the first week of class, students were asked to rank their prior experience to gene expression analysis, analysis of large data sets, and using R or other programming tools in data analysis and data visualization. Data shown are for 85 students in a genetics course who completed this survey in 2014. The actual number of students is designated for each answer choice.
| Activities | Assessment |
|---|---|
| Worksheet 1. Transcriptional Response to Cold Stress: Primary Literature Analysis and Developing Testable Hypotheses | |
| Observation and description: phenotypic effects of abiotic stress | Worksheet 1 (completeness and effort, feedback), lab report |
| Primary literature analysis: effects of abiotic stress on gene expression in plants | |
| Formulating hypotheses/predictions: number and types of genes affected by the stress and variation in response to different stress and between different genotypes | |
| Worksheet 2. RNA-seq Analysis: Principles | |
| Concept discussion: classes of RNA molecules, similarities and differences | Worksheet 2 (completeness and effort, feedback), lab report |
| Knowledge building: principles of RNA-seq analysis, creating libraries, and sequencing | |
| Worksheet 3. RNA-seq Analysis: Data Quality and Initial Analysis | |
| Understanding sequence read files (FastQ): how do my data look like? | Worksheet 3 (completeness and effort, feedback), lab report |
| Initial data analysis: data quality control using Green Line of the DNA Subway | |
| Analogy and exercise: principles of mapping and counting RNA-seq reads | |
| Worksheet 4. Data Analysis: Finding Differentially Expressed Genes | |
| DE-Seq analysis: finding differentially expressed (DE) genes | Lists of DE genes, summary tables, lab report |
| Formulating questions, choosing approaches to data visualization | |
| Data visualization and analysis | |
| Worksheet 5. Data Visualization: Common Types of Graphs Used to Show RNA-seq Data | |
| Exploring various approaches to RNA-seq data graphical visualization | Student presentations and discussion, worksheet 5, lab report |
| Data visualization and analysis | |
| Sharing the results with other groups, discussion of data and graphs | |
| Percent of correct answersb | ||
|---|---|---|
| Concept questiona | Pretest | Posttest |
| Genes and gene regulation (11 questions: 1–6, 8–11, 14) | ||
| Which of the following human cells contains a gene that specifies eye color? | 34 | 85 |
| In what way is the same environmental signal expected to modify gene activity in different individuals? | 19 | 71 |
| What proportion of genes is likely change their expression levels in response to environmental stress? | 13 | 87 |
| RNA-seq analysis (9 questions: 7, 12, 13, 15–18, 20, 21) | ||
| What is not true about RNA molecules that are “sequenced” during RNA-seq experiments? | 12 | 78 |
| What is not necessary to have in order to perform an RNA-seq experiment? | 68 | 86 |
| Data visualization (2 questions: 19, 22) | ||
| Two graphs below show the comparison of normalized gene counts from an RNA-seq experiment. What can you conclude based on these graphs? | 39 | 90 |
| Average score | |||
|---|---|---|---|
| Course and year | Number of students | Pretest | Posttest |
| Principles of Genetics, 2014 | 85 | 27 ± 15% | 79 ± 8% |
| Applied Biotechnology, 2015 | 8 | 52 ± 19% | 90 ± 10% |
Lab Implementation
The workflow of the RNA-seq analysis and activities performed by the students are shown in Figure 1. The complete list of student activities and associated assessment tools are shown in Table 1. All of the teaching materials are accessible under the Supplemental Materials.
Week 1.
Students completed three worksheets aimed at understanding the experimental system and an RNA-seq approach to investigating gene expression. They also developed interesting experimental questions and testable hypotheses related to the effects of abiotic stress on gene expression.
Worksheet 1: Transcriptional Response to Cold Stress: Primary Literature Analysis and Developing Testable Hypotheses. Students observed and described phenotypic effects of cold and heat stress on maize seedlings. They also conducted primary literature searches and briefly summarized two manuscripts describing the effects of exposure to abiotic stress in any plant system. Students were asked to use available information to predict the proportion and function of genes expected to respond to abiotic stress conditions in maize seedlings and to compare transcriptome response between different abiotic stresses (cold and heat) and in plants from different genetic backgrounds (B73 and Mo17). After a discussion of the major concepts of environmental effects on gene expression and an introduction of the RNA-seq data set, students formulated hypotheses regarding gene expression changes that could be answered using this data set.
Worksheet 2: RNA-seq Analysis: Principles. Students discussed the similarities and differences of major classes of RNA molecules and the means of separating mRNA from other RNA types and converting mRNA to DNA. They also investigated general approaches of Illumina RNA sequencing: fragmentation, adaptor ligation, indexing and multiplexing, and sequencing by synthesis. Using many available resources, including the Internet, textbooks, and help from the instructor, students constructed the schematic representation of the RNA-seq experimental flow and briefly described it in their own words.
Worksheet 3: RNA-seq Analysis: Data Quality and Initial Analysis. Students analyzed sample FastQ files, the output of RNA-seq experiments, to understand the format and content of the data produced by RNA-seq. Students also used the Green Line of the DNA Subway portal developed by the iPlant Collaborative (Goff et al., 2011; http://dnasubway.iplantcollaborative.org) to perform an initial analysis of the data quality for all abiotic stress maize samples (instructor-created public project “Maize Abiotic Stress”). Additionally, they discussed the ways data quality is graphically visualized in the DNA Subway software. Finally, students completed a short exercise demonstrating the principles of following steps of the RNA-seq analysis: mapping short reads back to the genome and read counting and normalization. We chose to work with the Green Line of DNA Subway, because it provides the intuitive platform for conducting some of the analysis, essential for students who lack computer programming skills and for a lab environment in which computer power and time are limited. An instructor-created public project (“Maize Abiotic Stress,” DNA Subway Green Line), which students could access from their computers, pre-ran quality-control analysis (Supplemental Figure 1; FastQC; http://dnasubway.iplantcollaborative.org). Although the DNA Subway Green Line allows the complete workflow of the RNA-seq analysis (Tuxedo protocol) to be conducted, many of the analysis steps take a long time and could not be completed in 3-h lab periods. Instead, the students were provided with the files containing raw counts for reads corresponding to all maize genes for all genotype/condition combinations, essentially “skipping” tedious steps of read mapping and counting. This approach allowed students to focus on principles of read alignment and counting through a series of guided exercises in worksheet 3 and on discovering differentially expressed genes in worksheet 4.
Weeks 2 and 3.
Students worked with files containing raw gene counts for two abiotic stresses and control samples for two genotypes generated by the instructor (see Data Set 1 in the Supplemental Material). The students identified questions of interest and created lists of differentially expressed genes for conditions relevant to their questions. They also discussed several approaches to visualizing RNA-seq data and used these approaches to answer the questions generated during the previous steps. Students informally presented their work to peers and the instructor to receive feedback and solve problems during the analysis.
Worksheet 4: Data Analysis: Finding Differentially Expressed Genes. This worksheet guides students through DE-Seq analysis in R statistical analysis software (Anders and Huber, 2010) and provides necessary explanations of the steps involved. Students performed data normalization and statistical analysis of differentially expressed genes and filtered their results based on the significance level, fold difference of expression levels, and the minimal expression level in one or several samples (see Data Set 2 in the Supplemental Material for an example of a DE-Seq output file). The whole class engaged in the discussion of criteria that should be used to identify genes as differentially expressed. The students discussed and chose the questions they would like to address and approaches to data visualization and analysis that could be used to answer their questions.
Worksheet 5: Data Visualization: Common Types of Graphs Used to Show RNA-seq Data. This worksheet aims at introducing students to various types of graphical representation of the RNA-seq data, such as scatter plots, histograms, kernel-density plots, heat maps, Venn diagrams, and genome views. It uses examples of figures from published RNA-seq studies and asks students to interpret these graphs. In addition to worksheet 5, students used a document, “How to Make Graphs in R” (see the Supplemental Material), to guide them through building graphs for visualizing their data.
Lab Assessment
Students’ experience of engaging with large-scale data analysis, gene expression, and RNA-seq analysis, as well as using R and other computational tools for analysis of biological data sets, was assessed during the first week of the course using a short survey (Supplemental Material). Students’ learning was assessed with a content assessment test, a set of 22 multiple-choice questions targeting general concepts of eukaryotic gene expression regulation as well as the principles of RNA-seq analysis and data visualization and interpretation (Table 2 and Supplemental Material). To assess student learning gains, we used the same test as a pretest and a posttest. Student scores were used to calculate normalized learning gains (Hake, 1998), a metric that takes into account differences in student knowledge and measures the fraction of the available improvement that can be gained. In addition, students were asked to complete a CURE survey (Lopatto et al., 2008) to assess students’ perception of their learning and development as scientists. A CURE presurvey and a content assessment pretest were conducted during week 2 of both courses, while a CURE postsurvey and a content assessment posttest were conducted during week 14, the last week of the courses, at least 1 wk after the lab reports were turned in. Owing to a low number of students in Applied Biotechnology, the assessment data described here refer to the students from Principles of Genetics, unless noted otherwise. Extra-credit points were assigned for correct answers to the content assessment pre- and posttests and for completion of CURE surveys. To assess student skills in data visualization and interpretation of graphs related to RNA-seq analysis, we assessed the results sections of the students’ lab reports using a rubric focusing on the appropriateness, clarity, and quality of the figures and figure legends and the interpretation of the data presented in the figures (see the Supplemental Material for the rubric used). In addition to the pre/posttest assessment, all of the student group worksheets were graded by the instructor, and all mistakes and misconceptions were discussed in class. Finally, students were asked to provide any unsolicited comments about the RNA-seq laboratory series as a part of the university-wide postcourse online student evaluations. These comments remained completely anonymous and confidential.
Plant Growth and Stress Conditions
B73 and Mo17 maize seedlings were grown at 24°C in 1:1 mix of autoclaved field soil and MetroMix under natural light conditions. For cold stress, seedlings were incubated at 5°C for 16 h. For heat stress, seedlings were incubated at 50°C for 4 h. Light conditions were the same for all stress and control conditions.
Data Set Description and Data Analysis
The RNA-seq data set of SRA Project PRJNA244661 was used in implementation of the lab exercises (Makarevitch et al., 2015). This data set includes three replicates of RNA-seq data from 14-d-old maize seedlings of two inbred lines, B73 and Mo17, grown under controlled conditions and subjected to cold and heat stress as described above (for the details of plant growth, sample collection, RNA isolation, library preparation, and sequencing, see Makarevitch et al., 2015). Transcript abundance was calculated by mapping reads to the combined transcript models of the maize reference genome (AGPv2) using TopHat (Trapnell et al., 2009). Reads were filtered to allow for only uniquely mapped reads. A high degree of correlation between replicates was observed (r > 0.98). RPKM (reads per kilobase of transcript per million reads mapped) values were developed using BAM to Counts across the exon space of the maize genome reference working gene set (ZmB73_5a) within the iPlant Discovery Environment (www.iplantcollaborative.org). Genes were considered to be expressed if RPKM > 1 and differentially expressed if log2(stress/control) > 1 or log2(stress/control) < −1. Statistical significance of expression differences was determined using the DE-Seq package (Anders and Huber, 2010). Gene ontology analysis was performed using information from the Maize Genetics Database (maizegdb.org).
RESULTS
Plant Materials and the RNA-seq Data Set
The key to successful implementation of this series of lab exercises is the choice of the data set for analysis. We chose a data set representing a transcriptional response to two different abiotic stress conditions, heat and cold, in two maize genetic backgrounds, Mo17 and B73 (Makarevitch et al., 2015). This data set allowed students to ask a variety of questions about the effects of abiotic stress on gene expression and offered a wealth of hypotheses that students could test. To provide students with the background on abiotic stress and experimental flow (Figure 1), we had students reproduce the conditions of the experiment and observe the effects of cold- and heat-stress exposure on maize seedlings (Figure 3). When the stressed plants were allowed to recover after stress for 24 h, phenotypic consequences became apparent for both stress treatments. While Mo17 plants were resistant to cold stress and showed very little, if any, phenotypic differences compared with control plants, B73 seedlings showed striking phenotypic response with dry and necrotic leaf edges and tips and severe wilting. Both Mo17 and B73 seedlings showed mild response to heat stress, with wilted and discolored leaves (Figure 3).
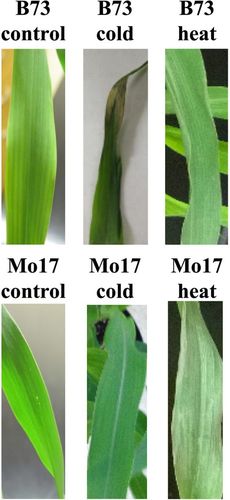

Figure 3. Phenotypic effects of exposure to abiotic stress observed by students. B73 seedlings show strong response to cold stress with dry necrotic leaf edges and tips, while Mo17 seedlings show only minimal response to cold. Both B73 and Mo17 seedlings show response to heat stress with wilted leaves.
Experimental Questions and Data-Visualization Approaches Chosen by Students
Guided by worksheet 1, students investigated primary literature on stress response in plants and formulated a series of questions that could be asked about the data (see Supplemental Table 1 for a list of students’ questions). The questions ranged from “Expression of how many genes is affected by cold?” to “What biochemical pathways are activated in response to stress?” Given variation in response to stress between maize seedlings of different genetic backgrounds, many students were interested in comparing the lists of genes affected by cold stress in Mo17 and B73 genotypes and in finding potential candidate genes that would explain the resistance of Mo17 to cold. Several student groups were interested in comparing genes that responded to different stress conditions, asking, “Do different abiotic stress conditions elicit similar or different responses in gene expression?” Students successfully ran DE-Seq analysis of the samples pertinent to their research questions and identified genes differentially expressed in response to abiotic stress. Students from different groups compiled a table summarizing the number of genes differentially expressed in response to different abiotic stress conditions in both genotypes (Supplemental Table 2). One of the most interesting discussions driven by students revolved around what genes should be called “differentially expressed,” the criteria that should be used to define “differentially expressed genes,” and whether these criteria should be uniform for a group of scientists working on the same problem. Generating lists of differentially expressed genes stimulated further questions. With some guidance, students explored the approaches to visualizing data and asked deeper questions about differentially expressed genes (worksheet 5). Students used a variety of approaches to visualize the data pertinent to their research questions (Supplemental Table 1 and Figure 4). Some groups investigated the level of individual variation in transcriptional response to abiotic stress by comparing variation between replicates of the same condition and between different samples using scatter plots (Figure 4, A and B). Other students asked the same question by constructing a heat map that visualized differentially expressed genes in two genotypes under stress conditions (Figure 4E). Several student groups compared the stress response between maize genotypes (Figures 4, C and E). Students also asked whether some genes responded in a similar manner to different abiotic stress conditions (Figure 4F). Finally, students investigated the likely functions of the stress-response genes by comparing the proportion of genes that belong to different gene ontology categories for all maize genes and genes differentially expressed in response to stress (Figure 4D).
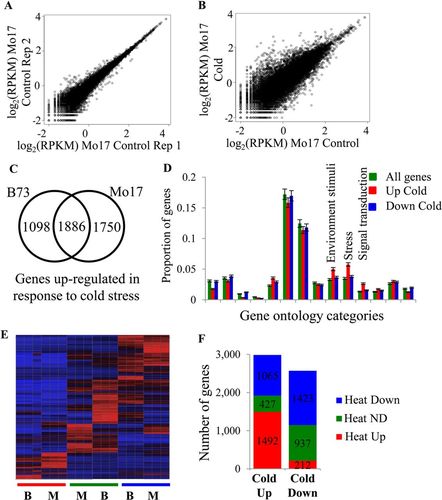
Figure 4. Examples of graphs students used to visualize data and answer the questions. (A and B) Comparison of variation between two replicates of the same condition and between stress and control conditions. Log2RPKM values are graphed for all maize genes. (C) The conservation of stress response. Many genes up-regulated in response to cold stress in B73 are also up-regulated in response to cold stress in Mo17, while many genes show response in only one of the genotypes. (D) The proportion of all maize genes, genes up-regulated in response to cold, and genes down-regulated in response to cold is shown. SE is shown with error bars. Three gene ontology categories significantly overrepresented among genes up-regulated in response to cold stress are shown (p < 0.05). (E) Abiotic stress exposure results in up- or down-regulation for a number of maize genes in each genotype. The Z-normalized RPKM values for all differentially expressed genes were used to perform hierarchical clustering of the gene expression values. The genotypes (B73: B; Mo17: M) and conditions (heat: red; control: green; cold: blue) are indicated below each column. Three replicates of each condition are shown. (F) Genes affected by cold stress are frequently up-regulated in response to heat stress as well. Genes up- and down-regulated for cold stress in B73 are shown, as is their response to heat stress. ND: the genes with no differential expression. The number of genes in each category is shown.
Assessment of Student Learning
A combination of subjective and objective assessment approaches were used to assess student learning as the result of this lab series. First, students were asked to complete a test with 22 multiple-choice questions once during the first week of the class (pretest) and once at the end of the last lab period (posttest). The proportion of correct answers increased from 27 to 79% (normalized learning gain of 0.71) for students in Principle of Genetics and from 52 to 87% (normalized learning gain of 0.73) for students in Applied Biotechnology (Figure 5 and Tables 2 and 3). Although there are no established criteria for what constitutes acceptable learning gains on these tests, a normalized gain of ≥0.50 probably represents a substantial achievement. The questions were designed to test understanding of principles of gene expression regulation, major concepts of RNA-seq analysis, and data analysis skills (see Table 2 for question category assignment). Although students were expected to be more familiar with gene expression regulation concepts compared with principles of RNA-seq analysis, average pretest scores for both categories were low (25 and 27% for RNA-seq analysis and gene expression regulation, respectively), possibly suggesting low emphasis on these topics in high school biology courses. Interestingly, the most difficult questions from the regulation of gene expression category (questions 3, 9, and 14) focused on the overall transcriptional response to stress and its magnitude and variation. The overwhelming majority of students in Principles of Genetics said that stress affects gene expression in a predictable way, primarily activating gene expression of a relatively small number of genes. Conversations with the students during their work on primary literature analysis and, especially, during their analysis of differentially expressed genes, confirmed these observations, since students were very surprised to see that as many as 10% of maize genes could be either up- or down-regulated in response to stress with response varying between maize seedlings of different genetic background.
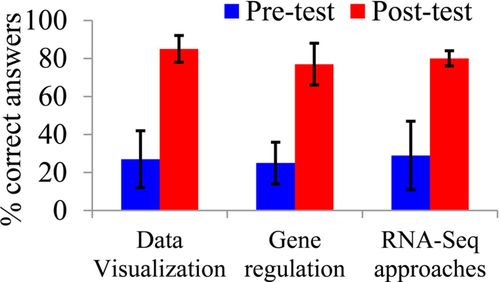
Figure 5. Assessment of student learning. Student learning was assessed using a test consisting of 22 multiple-choice questions. Questions were separated into three categories, and the average proportion of correct answers for the questions in these categories was calculated for two courses (Principles of Genetics and Applied Biotechnology). Vertical bars show SD. For all three categories, the differences between pretest and posttest were significant as tested by paired t test (p < 0.01).
For assessment of student skills in graphical data visualization and interpretation, 27 group lab reports were assessed using the rubric that focused on the appropriateness, clarity, and quality of the figures, figure legends, and data interpretations (Table 4; see the Supplemental Material for the rubric used). Only three of 27 lab reports (11%) failed to achieve the level of “accomplished” (15/20 points), while seven reports (26%) scored 19 or 20 points. Average scores in all five rubric categories exceeded the level of “accomplished” (3/4 points), demonstrating that the students were able to state appropriate experimental questions, choose and build adequate data visualizations, and interpret the results of their experiments.
| Rubric category | Criteria for the correct responses | Student scores (out of 4 points for each category) |
|---|---|---|
| Experimental question | Clarity and appropriateness of the experimental question | 3.46 ± 0.68 |
| Graphs | The choice of the visualization approach and the correct organization of the graph | 3.20 ± 0.62 |
| Graph labels | Presence and accuracy of the graph labels | 3.42 ± 0.60 |
| Figure legends | Completeness and accuracy of the figure legends | 3.25 ± 0.64 |
| Data interpretation | Clarity and appropriateness of the conclusions, support of the conclusions by the graphs | 3.20 ± 0.70 |
| Total | 16.45 ± 2.44 |
For assessment of student perception of the lab series and the learning gains, a CURE survey was implemented (Lopatto et al., 2008). Students reported perceived learning gains higher or comparable with learning gains reported by all CURE participants in all 21 categories, with the largest gains in categories related to understanding the scientific process and skills in data analysis (Table 5). Finally, students were asked to provide comments regarding the RNA-seq data analysis lab experience in the anonymous university-wide online student evaluations of the course, and 65 students chose to provide comments. All student responses were analyzed using the constant comparative method (Erickson, 2012). Student comments were initially coded using open codes, such as “challenging,” “frustrating,” “engaging,” “exciting,” “real research,” “large data,” “real tools,” and “confusing.” Initial codes were combined into conceptual codes that were used to identify the themes emerging from the data. Emerging themes identified in the analysis characterized the students’ engagement and perceptions of the laboratory series on RNA-seq analysis as a difficult and engaging real research experience in computational biology: “exciting and interesting,” “authentic research,” “computational nature of biology research,” and “discontent and frustration” (Table 6). While most of the students reacted positively to the experiences of this laboratory module, greater than 20% of the students included comments suggesting that the activities were too complex or some of the aspects of the analysis were difficult to complete during the time allotted. We believe that extending this laboratory module to four or even five lab periods by incorporating additional debriefing activities, or even minilectures provided by the instructor and aimed at explaining most common mistakes, would significantly ameliorate this problem.
| Category | Genetics learning gains (65 students) | CURE participants (4800 students) |
|---|---|---|
| Understanding science process | ||
| Understanding how knowledge is constructed | 3.49 | 3.42 |
| Understanding the research process | 3.50 | 3.46 |
| Understanding how scientists work on real problems | 3.62 | 3.58 |
| Understanding that scientific assertions require supporting evidence | 3.59 | 3.64 |
| Understanding science | 3.66 | 3.58 |
| Data analysis skills | ||
| Ability to integrate theory and practice | 3.38 | 3.46 |
| Ability to analyze data and other information | 3.96 | 3.74 |
| Skill in interpretation of results | 3.62 | 3.54 |
| Ability to read and understand primary literature | 3.45 | 3.34 |
| Communication skills | ||
| Skill in science writing | 3.39 | 3.31 |
| Open codes | Theme | Description | Student quotes |
|---|---|---|---|
| Cool lab | Exciting and interesting | Overall perception of the lab series | “I never had so much fun building graphs.” |
| Interesting | “Great addition to Genetics.” | ||
| Unusual lab | “The lab was very frustrating and difficult, but I learned a lot!” | ||
| Fun | |||
| Real research | Authentic research | Includes references to the research nature of the lab series | “Doing real research in class is really cool.” |
| Real science | “We worked with real data on real research problem[s].” | ||
| Cool experiment | “Nobody knew the answers to our questions.” | ||
| Real data | “We got to build graphs in R and they looked like the graphs from the papers we were reading!” | ||
| Programming | Computational nature of biology research | Describes the student perception of programming and computational studies as a part of biology | “This was the first time I was involved in large data analysis; it would be great to do it more often!” |
| Bioinformatics | “I never realized that biology is almost computer science now.” | ||
| Databases | “I wish I knew more programming and was more familiar with computers, this was fun!” | ||
| A lot of computation | |||
| Large data sets | |||
| Confusion | Discontent and frustration | Reflects negative perceptions of the lab series due to lack of interest, confusion, or frustration | “This lab is way too difficult and should not be a part of introductory course.” |
| Frustration | “I was confused through the whole three weeks.” | ||
| Analysis did not work | “My R code never worked and the instructor had to fix it all the time. Very frustrating.” | ||
| Lack of engagement | |||
| Too complex |
DISCUSSION
We developed a series of laboratory exercises that engages students in investigating transcriptional response of maize seedlings to abiotic stress. In our experience, a connection to climate change served as a great way to excite students about plant genetics and show them the relevance of plant genetics research. Analysis of student comments in the online course evaluations suggests that students were excited to participate in the real research project and analyze the unpublished data, potentially exploring novel scientific ideas and connections (Table 6), highlighting the need for a careful choice of the RNA-seq data set. We chose a data set that was not fully characterized by the time of the lab implementation and plan to develop a novel data set for the next year’s course based on the ideas the students developed in Principles of Genetics. One of the main advantages of the approach we used is the opportunity to engage students using any publicly available RNA-seq data set. Since its introduction, RNA-seq, the tool to precisely measure the levels of transcripts, has revolutionized our view of the extent and complexity of eukaryotic transcriptomes (Wang et al., 2009). The SRA of the NCBI is a public repository containing more than 150,000 RNA-seq data sets that are freely available for download. The manuscripts describing these data sets usually address specific questions and leave a lot of room for additional questions that students could investigate. Furthermore, the costs for library construction and sequencing, the most expensive steps of generating RNA-seq data, continue to decrease, and the possibility of running RNA-seq experiments designed and run by students in undergraduate biology courses is already within reach for many institutions. Most of the exercises and the general approach described here can be easily adopted for analysis of any RNA-seq data set. The series of laboratory exercises on transcriptional response to abiotic stress in maize was implemented in the introductory genetics course and in the upper-level biotechnology course and as an approach to introduce summer research students to RNA-seq analysis. Depending on time commitment and the level of the students, these exercises could be extended to incorporate quantitative reverse-transcription polymerase chain reaction (qRT-PCR) validation of most interesting differentially expressed genes as well as to test expression of these genes under other relevant conditions, further investigating the biological role of identified differentially expressed genes. Approaches to integrating qRT-PCR, as well as primer design, into undergraduate lab exercises have been previously described (Robertson and Phillips, 2008; Hancock et al., 2010).
One of the difficulties in incorporating RNA-seq analysis into the classroom is the complexity of the tools used by the research community to map and count RNA-seq reads and to find differentially expressed genes. The DNA Learning Center in collaboration with iPlant developed a Green Line on the DNA Subway website that allows for storage and analysis of the RNA-seq data. Many features of the Green Line are readily accessible, and the students were able to conduct analyses and interpret their data. Unfortunately, the time required for running the applications for read mapping and counting for a large maize genome on the Green Line was too long to be effectively integrated in a time-limited lab environment. To overcome this issue and to allow students to concentrate on data analysis instead of technical details of the computer applications, we chose to provide students with the raw read counts. Students were engaged in a series of exercises simulating these activities, including analysis of analogies aimed at helping them understand the purpose and potential limitations of each of the steps. Students used a DE-Seq R package to find differentially expressed genes and to conduct downstream analysis of these genes (Anders and Huber, 2010). While the students were provided with the template scripts for DE-Seq analysis and using R to build various graphs, students had to modify these scripts to their specific questions, a task that required them to understand the purpose of each line of code. Such an approach allowed avoidance of some of the apprehension toward programming and incorporated genuine biologically relevant programming experience that went beyond the use of “black box software” as called for by BIO2010 (NRC, 2003). This lab series introduced many mathematical skills, including data normalization and statistical testing of differential gene expression, through real-world examples, an approach shown to result in higher learning gains in quantitative reasoning skills for biology students (Matthews et al., 2010; Feser et al., 2013; Hester et al., 2014). In addition, the instructional approach described here, specifically peer-to-peer presentations and peer reviews of the lab reports, presents potential for students to develop written and oral communication skills. Although beyond the scope of this project, formal assessment of development of mathematical and communication skills as the result of implementing this laboratory series should provide interesting data on integrative development of student skills related to science.
In addition to teaching concepts of gene expression and regulation in response to changes in environmental conditions, this laboratory series aimed to increase student skills in data analysis. One of the major emphases of our approach was to help students analyze various types of graphs that are common in primary literature describing RNA-seq data and to provide students with the opportunities to build similar graphs using their own data. In framing graph analysis exercises in worksheet 5, we enhanced many of the ideas from Figure Facts (Round and Campbell, 2013) with the peer-to-peer presentations of the primary literature graphs and found this approach to be very effective. An opportunity to build and present graphs similar to the ones seen in primary literature using students’ own data further enriched this experience in data visualization. Such an approach of mimicking the peer-review process used by scientists through critiquing one another’s papers has been demonstrated to be beneficial for students (Guilford, 2001). Our objective assessment data (relevant questions in the test and the quality of figures and data interpretation in the students’ reports) and subjective assessment data (the CURE survey) suggest that engaging in the RNA-seq analysis laboratory experience led to learning gains in data analysis and interpretation skills.
The National Science Education Standards and education research literature emphasize that students need to develop skills in quantitative data analysis (NRC, 2003; Bialek and Botstein, 2004; AAAS, 2011; Feser et al., 2013). Biology undergraduate students are lacking opportunities to be directly involved in quantitative data analysis, especially in analysis of large data sets that have become a “staple food” of current biology research. In addition, biology students lack mathematical and computational skills necessary for data analysis and perceive mathematics as irrelevant to their field (Zan et al., 2006). This problem is well recognized by the community, and a concerted effort to infuse computational and mathematical training into biology courses will likely help in developing more opportunities for students to develop these skills (Caudill et al., 2010; Milton et al., 2010; Sorgo, 2010; Feser et al., 2013; Hester et al., 2014). As measured by the Survey of Undergraduate Research Experiences and the CURE survey, authentic undergraduate research experiences provide significantly higher gains in data analysis skills, as well as in many other areas, including knowledge and understanding of science and the research process, problem solving, communication skills, and critical thinking (Lopatto et al., 2008, 2014). Developing authentic student research experiences that incorporate large data analysis is hindered by the high level of complexity impeding students’ ability to fully comprehend the problem and by limited access to the computational tools and data sets. In addition, projects should provide students with opportunities to develop independent research questions and should be engaging for students and relevant to the course in which they are embedded. RNA-seq experiments are particularly flexible in that regard. RNA-seq analysis is widely used in research projects across most fields of biology and across most biological systems, making it a great tool to excite students interested in different biological processes and providing ample data to allow students to investigate independent research questions. Incorporating RNA-seq analysis in a biology curriculum provides unique opportunities to involve students in real biological research, improving students’ skills in data analysis, data visualization, and science communication.
ACKNOWLEDGMENTS
The authors are grateful to Peter Hermanson, Amanda Waters, Kathryn Malody, Hailey Karlovich, Josie Slater, Amanda Nimis, and Kristin Male for help and support in developing protocols for stress conditions, collecting samples, and validating data; Nathan Springer for continuous support and encouragement; David Micklos, Mona Spector, and Judy Brusslan for helpful and inspirational discussions around undergraduate education and research; Erin Dolan for detailed and thoughtful suggestions for improving the manuscript; and all of the students in I.M.’s courses. This work was supported by National Science Foundation grants MRI-R2 0957312 and DIOS 1237931 to I.M.